Professor Lizhen Huang's Team Publishes in ACS Sensors : One-Step Extraction-Free RPA-Cas12 Technology Enables Accurate Differentiation of African Swine Fever Wild-Type and Deleted Strains
A research team led by Associate Professor Huang Lizhen from the School of Biology and Biological Engineering at South China University of Technology (SCUT) has published a study in ACS Sensors, a prestigious journal in the fields of biosensing and analytical chemistry. The paper is entitled “An Extraction-free One-Pot Assay for Rapid Field Discrimination of African Swine Fever Virus Variants by a Single-Step RPA-CRISPR/Cas12a Strategy”. Li Wenyan and Yang Yunpeng, 2022 master’s students at the School, are the co-first authors. Associate Professor Huang Lizhen, Hu Xianghua from Guangdong Medical Devices Quality Surveillance and Test Institute, and Li Xueping from Guangzhou Yoyoung bio-tech company serve as co-corresponding authors, with the School of Biology and Biological Engineering, South China University of Technology as the primary corresponding affiliation.Original link: https://doi.org/10.1021/acssensors.5c03287

In this study, we established the CORDSv2 one pot, one step detection system, which possesses the core advantages of nucleic acid extraction free, cold chain independent, ultra simple operation, high sensitivity, and high specificity. Precise genotyping of African swine fever virus (ASFV) strains can be accomplished within 40–60 minutes, completely breaking the site and operational limitations of conventional detection methods. This work provides an efficient and practical technical solution for frontline on site epidemic prevention at pig farms, slaughterhouses, and other field settings.
Currently, circulating ASFV strains in the field consist of highly lethal wild-type strains and clinically cryptic gene-deleted strains with mild symptoms. Conventional detection methods only confirm viral presence without strain differentiation; approaches targeting deletion sites are prone to false-negative results and incomplete coverage. Moreover, most technologies require laboratory-based nucleic acid extraction and are incompatible with rapid on-site detection at pig farms and slaughter points. This study was designed to address these critical challenges and achieve precise genotyping with minimal operational steps.
In this study, comprehensive phylogenetic and genomic analyses were performed on a large panel of ASFV strains. The results demonstrate that dominant gene-deleted strains predominantly belong to genotype II. Although deletion patterns vary considerably in the MGF, CD2V, I177L and other regions, the vast majority of gene-deleted strains harbor insertions of eGFP or mCherry reporter genes. This pivotal discovery revises the conventional detection paradigm: instead of targeting highly variable deletion regions, the assay focuses on stably conserved reporter genes and identifies gene-deleted strains via positive signals, yielding more reliable and unambiguous outcomes, and laying a solid scientific foundation for the entire detection system.
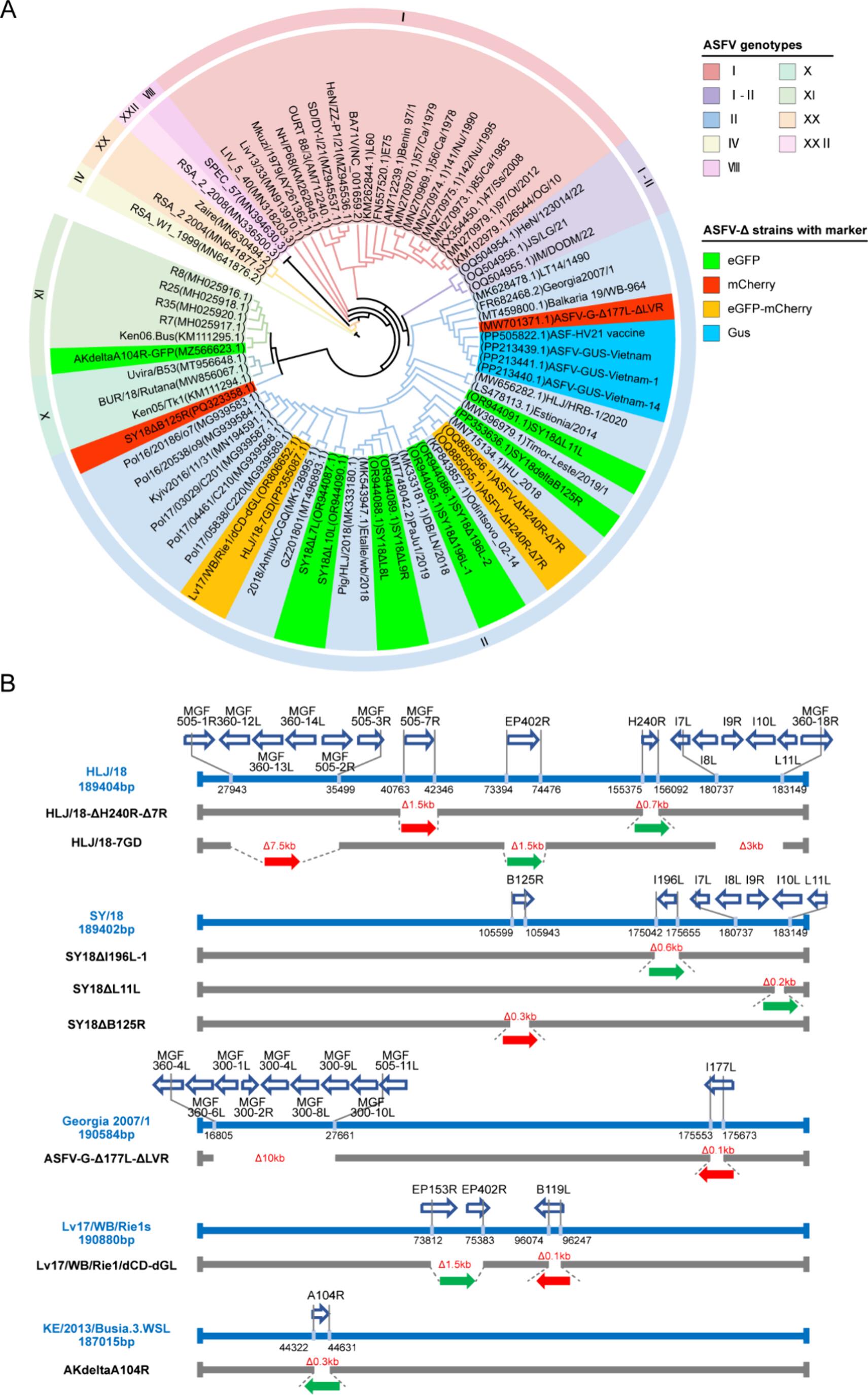
Figure 1. Phylogenetic and genomic characterization of wild-type and gene-deleted ASFV strains.
In this study, based on the self-developed AutoCORDSv2 platform, crRNAs and RPA primers were separately designed for eGFP and mCherry reporter genes, and the optimal detection combinations were obtained through multi-dimensional screening and systematic optimization. The eGFP-targeting system achieved a limit of detection (LOD) of 10-17 M with stable signal readout at 42 min; the mCherry-targeting system attained an LOD of 10-16 M with rapid readout at 30 min. Both systems exhibit outstanding sensitivity and specificity, with no cross reactivity against the swine genome or common swine pathogens, and deliver consistent and reliable results by either instrumental quantification or visual readout using portable devices. The dual-channel design enables comprehensive capture of gene-deleted strains carrying distinct reporter genes, resolving the limitations of incomplete coverage and ambiguous interpretation associated with conventional detection methods.
This study achieves three major breakthroughs for field deployment: extraction-free detection, wherein blood or saliva samples can be directly analyzed after 10 min of treatment with a high-efficiency lysis buffer; lyophilized microsphere reagents, which feature rapid reconstitution, excellent stability and elimination of cold-chain requirements for transport and storage; one-pot single-step operation, involving only three simple steps (sample addition, incubation, signal readout) accessible to non-specialist operators. When combined with a portable fluorescence detector, high-precision detection is truly translatable from the laboratory to grassroots frontline applications.
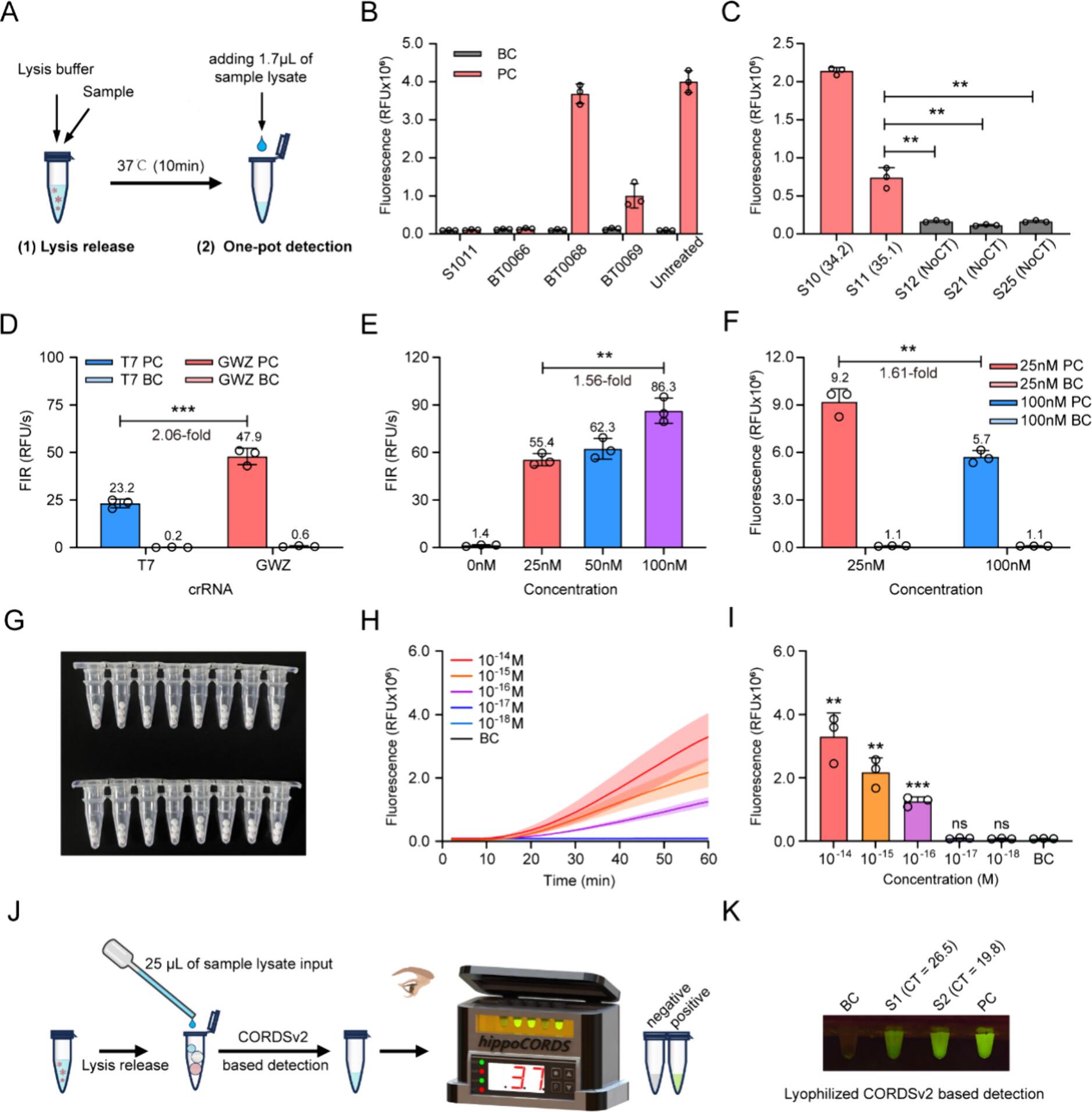
Figure 2.Development and clinical validation of an extraction-free and single-step lyophilized CORDSv2 system for field ASFV diagnosis.
The multiplexed CORDSv2-p72/eGFP/mCherry assay was constructed in this study: the conserved p72 gene was used to determine ASFV positivity, and reporter genes were employed to discriminate wild-type strains from gene-deleted strains. Validation using 96 clinical samples revealed that CORDSv2 exhibited an overall accuracy of 97.9% and a specificity of 100%, showing high consistency with the qPCR gold standard. The assay accurately identified wild-type strains and eGFP-labeled gene-deleted strains without false-positive signals or cross-reactions, completing the entire workflow within 60 min with an error rate below 3%, fully satisfying the requirements for on-site rapid screening and genotyping.
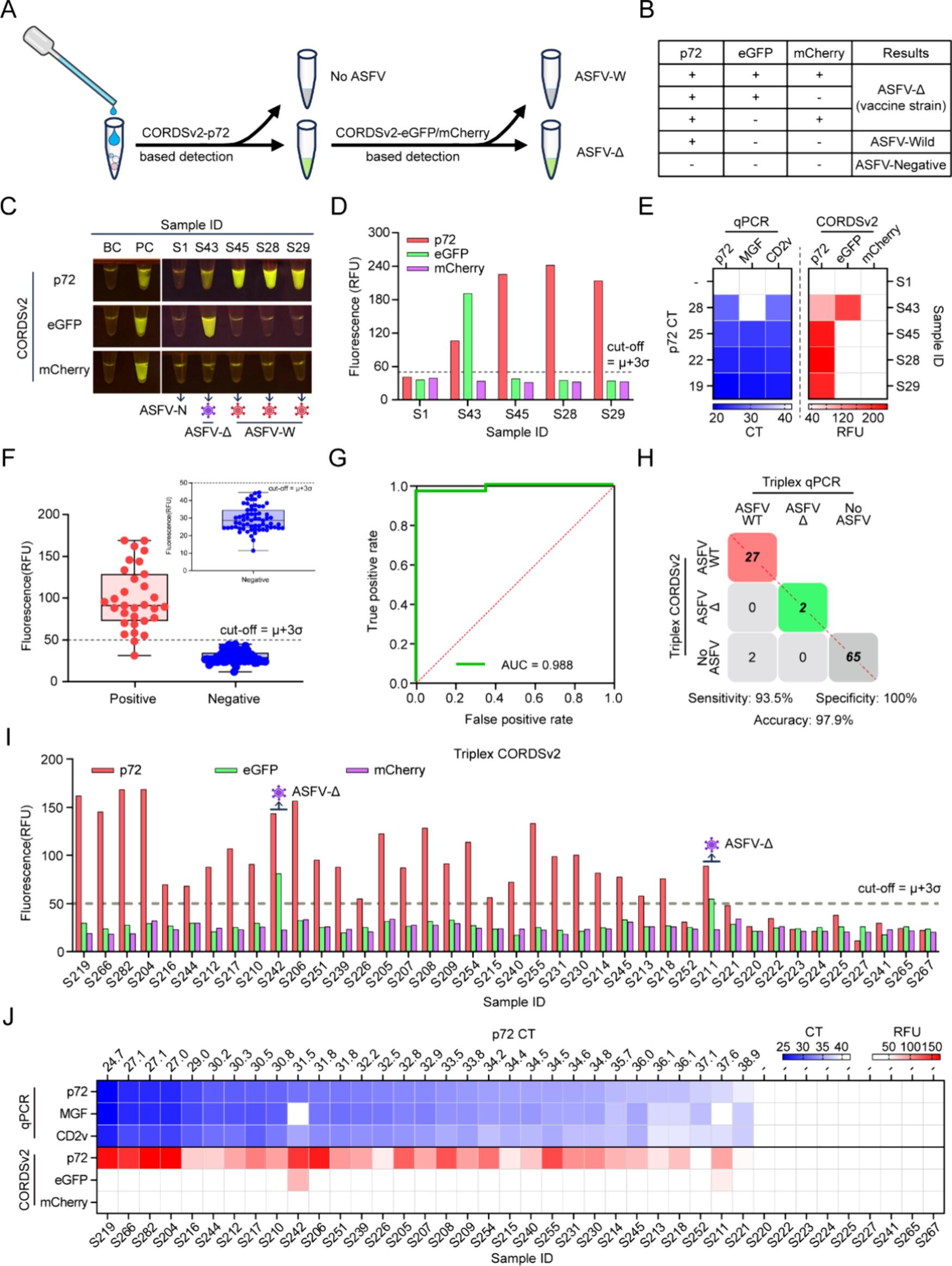
Figure 3. Clinical validation of triplex CORDSv2 assay for differential detection of ASFV wild-type and vaccine strains.
In summary, this study establishes a dedicated detection system targeting the eGFP/mCherry reporter genes unique to ASFV gene-deleted strains, enabling highly sensitive and specific genotyping of African swine fever. The technology also offers field-adapted advantages including extraction-free processing, lyophilized cold-chain-free reagents and one-pot single-step operation, accomplishing qualitative analysis and genotyping within 60 min with high clinical accuracy. This innovation effectively resolves the long-standing obstacles of conventional methods, including difficulties in on-site strain differentiation and high rates of missed detection, providing a simple, robust and reliable solution for frontline precise prevention and control of African swine fever.
This work was supported by the Natural Science Foundation of Guangdong Province (2022A1515011733, 2025A1515010459), the Guangdong Province Drug Administration Science and Technology Innovation Project (2024ZDZ07), the Science and Technology Program of Guangzhou (202206010073, 2023A03J0542), among others.