Huang Lizhen and Du Hongli’s joint team published in Nucleic Acids Research: IPGRM technology reveals genome-wide DNA repair network in CRISPR/Cas9 editing
A joint research team led by Associate Professor Lizhen Huang and Professor Hongli Du from the School of Biology and Biological Engineering at South China University of Technology (SCUT) has published a research article entitled “Indel pattern-guided repair mapping reveals genome-wide DNA repair networks in CRISPR/Cas9 editing” in the prestigious journal Nucleic Acids Research. Ph.D. student Yanbin Wan (Class of 2025), M.S. student Dongchao Huang (Class of 2021), Xuanye Zhao(Class of 2023) and Xiding Lin (Class of 2023) are co-authors of the study. Associate Professor Lizhen Huang and Professor Hongli Du serve as co-corresponding authors. The School of Biology and Biological Engineering, South China University of Technology, is the primary corresponding institution.
Article link: https://doi.org/10.1093/nar/gkag260.

CRISPR/Cas9 enables genome editing by introducing DNA double-strand breaks (DSB), and the final editing outcomes are largely determined by cellular DNA repair pathways. However, the molecular mechanisms and regulatory networks underlying different repair outcomes have not been systematically characterized, which limits the predictability and precise control of genome editing.
In this study, the team first performed high-throughput sequencing and systematic analysis of Cas9 editing outcomes. Despite their apparent randomness, the researchers identified highly structured and reproducible indel patterns. From a sequence perspective, templated 1 bp insertions dominate insertion events, while microhomology-mediated deletions are strongly enriched among deletion outcomes. Based on these observations, Cas9 repair outcomes were classified into seven distinct and recurrent patterns. From a kinetic perspective, these seven patterns exhibit characteristic temporal dynamics: 1 bp insertions arise rapidly at early stages, reflecting the fast repair kinetics of the non-homologous end joining (NHEJ) pathway; microhomology-mediated deletions accumulate gradually at later stages, indicating delayed activation of the microhomology-mediated end joining (MMEJ) pathway; meanwhile, large insertions and mutations show sustained accumulation over time. Together, these findings demonstrate that DNA repair is not a random process, but rather a temporally programmed and dynamic system involving both competition and coordination among distinct repair pathways.

Figure 1. Temporal dynamics of repair patterns across multiple genomic loci induced by Cas9
Building on these observations, the team developed an innovative screening strategy termed Indel Pattern-Guided Repair Mapping (IPGRM). The key methodological advance of IPGRM lies in encoding seven repair outcome categories—1 bp insertions (1 bp Ins), large insertions (Large Ins), non-microhomology small deletions (NonMH small Del), non-microhomology large deletions (NonMH Large Del), microhomology-mediated small deletions (MH small Del), microhomology-mediated large deletions (MH Large Del), and complex mutations (Mut)—into an automated sequence-based classification algorithm. This enables rapid, high-throughput identification and categorization of millions of editing events at single loci within seconds. This methodological breakthrough allows, for the first time, unbiased and systematic genome-wide screening of regulatory factors governing diverse DNA repair pathways underlying different editing outcomes. Compared with conventional approaches, which are typically limited to medium-throughput screens or a small subset of repair outcomes, IPGRM achieves genome-wide scale screening and simultaneous analysis of all seven repair categories using the same sequencing input, significantly enhancing both throughput and analytical resolution.

Figure 2. Schematic workflow of IPGRM classification
Leveraging the IPGRM platform, the research team constructed a genome-wide CRISPR screening library covering approximately 19,050 protein-coding genes and 1,864 miRNA genes, and performed comprehensive repair-seq analysis. This enabled systematic identification of key regulators associated with distinct indel patterns. Functional clustering analysis revealed three major regulatory modules: MMEJ-driven deletions form a coordinated regulatory module defined jointly by protein-coding genes and miRNAs, rather than being determined solely by core repair enzymes;NHEJ coordinates 1 bp insertions and non-microhomology deletions, with RFC4/5 stabilizing repair templates to suppress large deletions; atypical repair outcomes exhibit distinct genetic signatures, with large insertions associated with polymerase-related regulators and complex mutations enriched for chromatin-associated factors. Notably, S100A8 was identified as a potent suppressor of MMEJ through direct interaction with PARP1, revealing an underappreciated crosstalk between inflammatory signaling and DSB repair pathway choice.
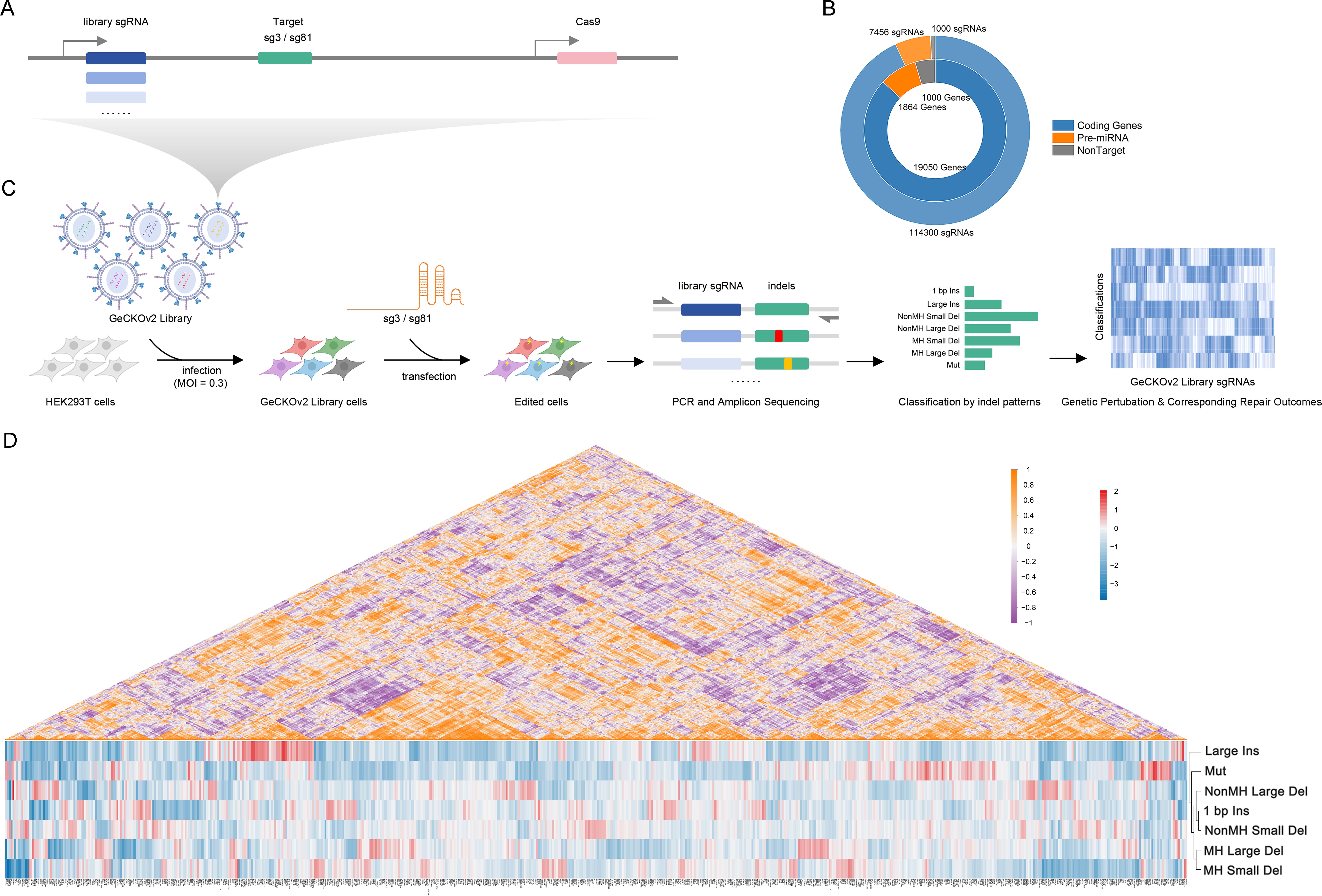
Figure 3. IPGRM identifies key regulatory factors involved in distinct repair patterns
Importantly, the study further demonstrates that loss of numerous miRNAs can significantly reshape the spectrum of Cas9 editing outcomes, with different miRNAs exhibiting distinct regulatory preferences. Some miRNAs selectively regulate microhomology-mediated deletions (MH small Del and/or MH large Del), others specifically influence 1 bp insertion events, while certain miRNAs simultaneously modulate the competitive balance among multiple repair pathways. These findings expand the regulatory landscape of DNA repair from protein-coding genes to non-coding RNA layers, highlighting the critical role of post-transcriptional regulation and opening new avenues for precise control of genome editing outcomes.
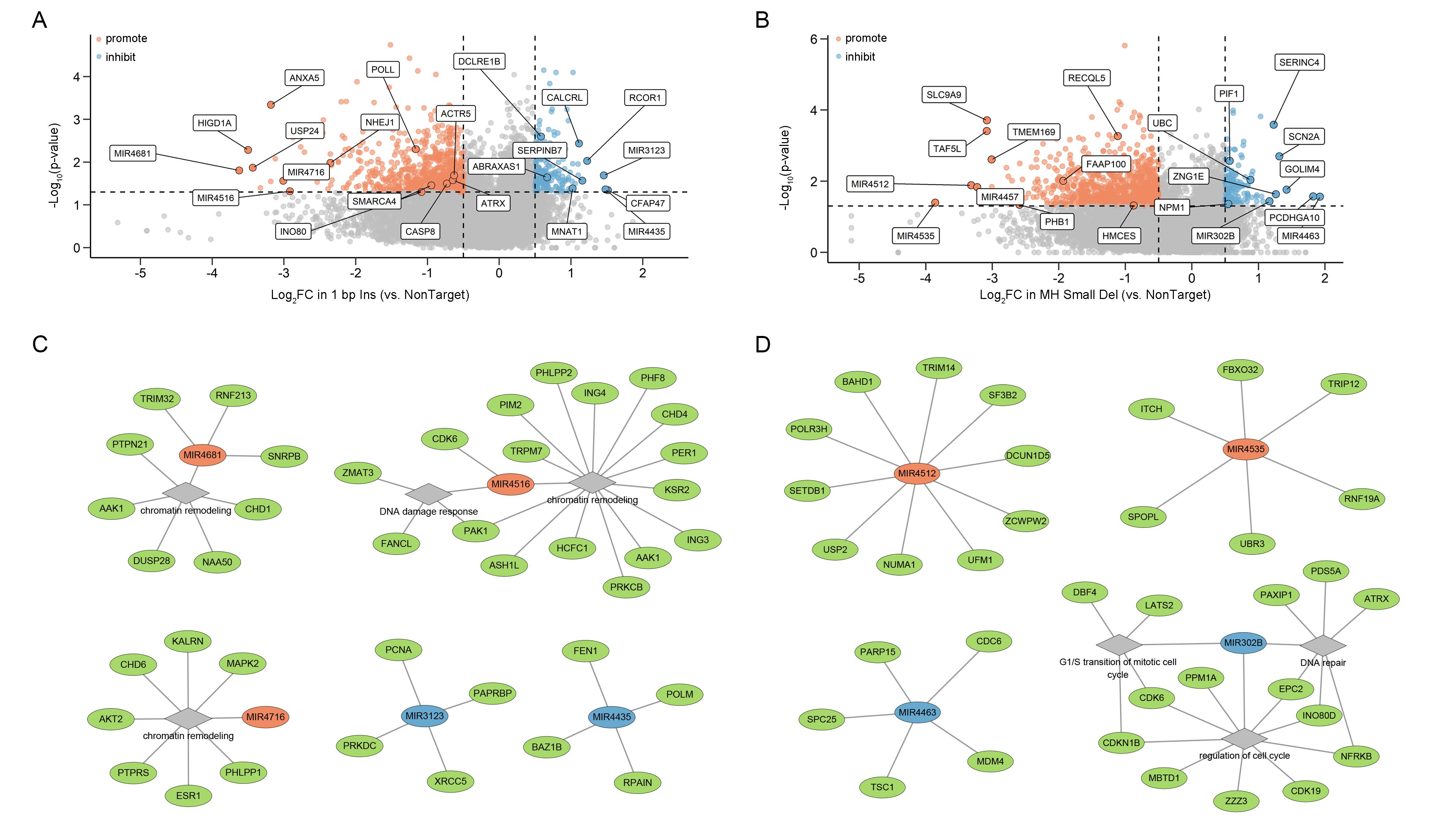
Figure 4. IPGRM reveals multilayered regulation of DSB repair by protein-coding genes and miRNAs
In summary, this study establishes the IPGRM framework based on indel patterns, systematically uncovers the temporal dynamics and genome-wide regulatory networks of DNA repair during CRISPR/Cas9 editing, and significantly advances our understanding of DNA repair mechanisms. These findings provide important theoretical support for improving genome editing precision and developing novel therapeutic strategies.
This work was supported by the Natural Science Foundation of Guangdong Province (2022A1515011733, 2025A1515010459), the Guangdong Province Drug Administration Science and Technology Innovation Project (2024ZDZ07), the Science and Technology Program of Guangzhou (202206010073, 2023A03J0542), among others.