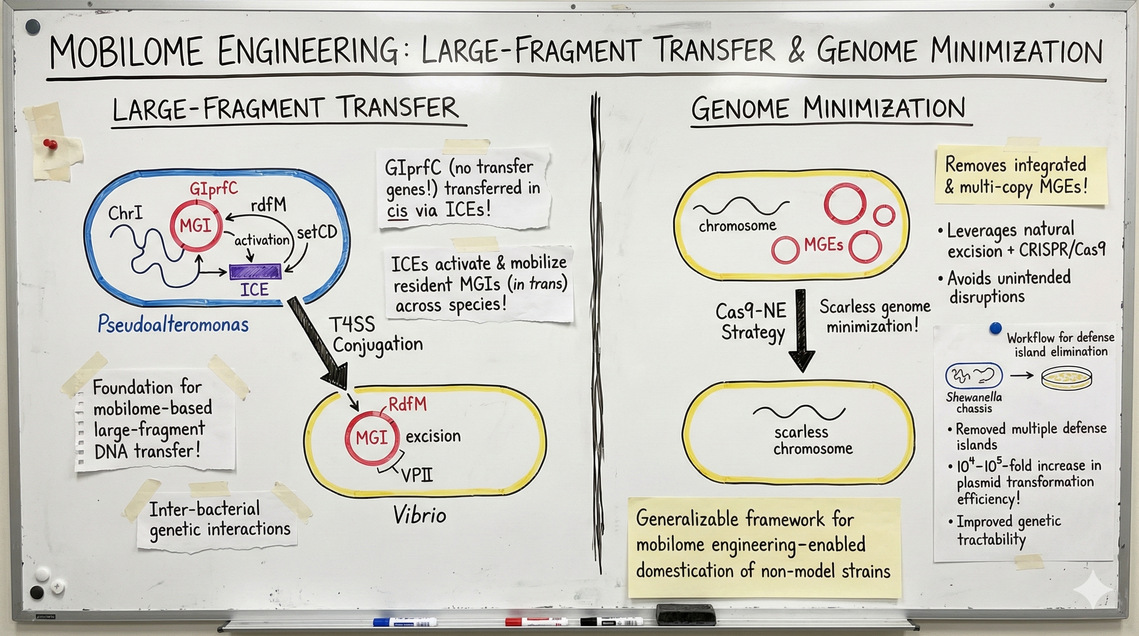
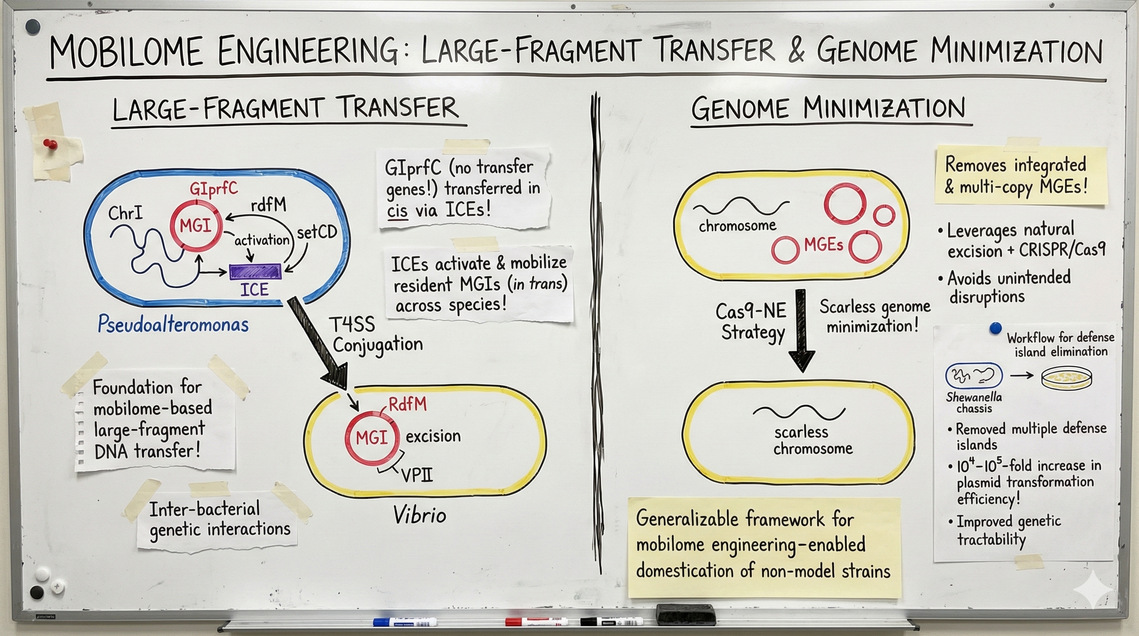
我们长期致力于探究细菌间遗传相互作用的机制。在假交替单胞菌研究中,我们发现基因组岛GIprfC虽缺乏典型的转移基因,却可通过与整合接合元件(ICEs)的顺式串联整合实现转移。进一步研究发现,ICEs能激活并驱动定植于基因组内的可移动基因组岛(MGIs),实现跨物种的反式转移。这些发现为基于移动基因组的大片段DNA转移机制奠定了理论基础。为实现精准的基因组精简,我们开发了Cas9-NE策略。该策略能同步清除整合态与多拷贝环状形式的移动遗传元件(MGEs),通过利用MGEs自然剪切机制结合CRISPR/Cas9靶向切割,有效避免了侧翼基因组区域的非预期破坏,实现了无缝基因组精简。此外,我们还建立了防御岛预测、定位与精准清除的全流程方法。以希瓦氏菌为模型底盘,成功移除多个防御岛后,其质粒转化效率提升10⁴–10⁵倍,显著增强了遗传可操作性。这一策略为通过移动基因组工程驯化非模式微生物菌株提供了普适性技术框架。
参考文献 https://journals.asm.org/doi/10.1128/aem.00095-24
参考文献 https://journals.asm.org/doi/10.1128/aem.00095-24
[en]We have long investigated the mechanisms underlying inter-bacterial genetic interactions. In Pseudoalteromonas, we discovered that the genomic island GIprfC, despite lacking canonical transfer genes, can be transferred in cis via tandem integration with integrative and conjugative elements (ICEs). Furthermore, we found that ICEs can activate and mobilize resident mobilizable genomic islands (MGIs), enabling their in trans transfer across species boundaries. These findings established a foundation for mobilome-based large-fragment DNA transfer. To achieve rational genome streamlining, we developed a Cas9-NE strategy capable of removing both integrated and multi-copy circular forms of mobile genetic elements (MGEs). This approach leverages the natural excision process of MGEs in combination with CRISPR/Cas9 targeting, thereby avoiding unintended disruptions of flanking genomic regions. This enables scarless genome minimization. In addition, we established a workflow for the prediction, localization, and precise elimination of defense islands. Using Shewanella as a model chassis, we successfully removed multiple defense islands, leading to a 10⁴–10⁵-fold increase in plasmid transformation efficiency and significantly improving its genetic tractability. This strategy provides a generalizable framework for mobilome engineering–enabled domestication of non-model microbial strains.
Refer to ISME J. 2022, 16(9), 2220-2229, https://www.nature.com/articles/s41396-022-01272-1
Appl Environ Microbiol. 2024,18:e0009524, https://journals.asm.org/doi/10.1128/aem.00095-24
[/en]