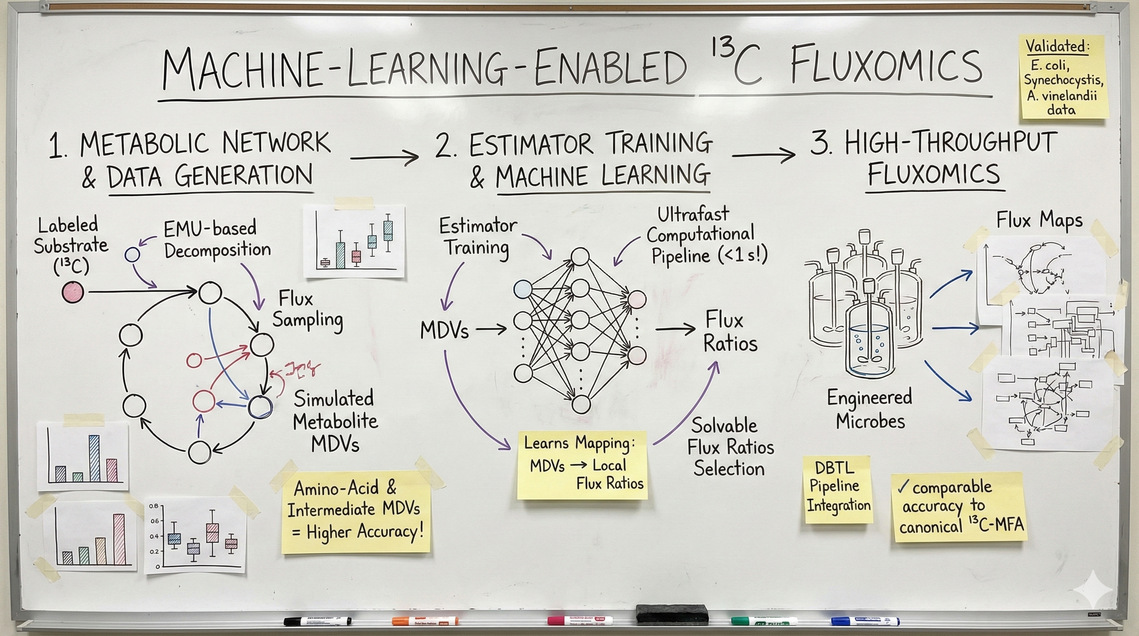
我们开发出了一种通用型超快速计算流程,该流程通过学习从模拟质谱同位素峰群分布向量(MDV)到局部通量比值的映射关系,进而结合实测交换通量解析细胞内全局通量分布。该工作流通过基于代谢物单元的网络分解与通量采样自动生成训练数据,程序化筛选通量比值的“可解性”,遴选信息丰富的MDV特征,并为每个通量比值训练专用估计器(深度神经网络与集成学习方法)——同时利用氨基酸及中间代谢物MDV数据可显著提升精度。经传统13C代谢通量分析在模拟及真实大肠杆菌数据上的验证,该方法在保持相当精度的同时,将预训练模型计算时间压缩至1秒以内,并内置适用于大肠杆菌、聚球藻PCC 6803及棕色固氮菌的即用模型,使之完全适配高通量代谢表型分析及设计-构建-测试-学习循环的集成应用。
参考文献 https://pubs.acs.org/doi/10.1021/acssynbio.1c00189
[en]We developed a generalizable, ultrafast computational pipeline that learns the mapping from simulated mass isotopomer distribution vectors (MDVs) to local flux ratios and then uses those predicted ratios together with measured exchange fluxes to solve global intracellular flux distributions. The workflow automatically generates training data by EMU-based metabolic network decomposition and flux sampling, programmatically screens flux-ratio “solvability”, selects informative MDV features, and trains per-ratio estimators (DNNs and ensemble methods) — using both amino-acid and intermediate MDVs substantially improves accuracy. Validated against canonical ¹³C-MFA on simulated and real E. coli data, the approach achieves comparable accuracy while shrinking compute time to <1 s with pretrained models, and includes ready models for E. coli, Synechocystis PCC 6803 and A. vinelandii — making it suitable for high-throughput metabolic phenotyping and integration into DBTL pipelines.

Refer to ACS Synth. Biol. 2022, 11 (1), 103−115, https://pubs.acs.org/doi/10.1021/acssynbio.1c00189 [/en]